library(fwildclusterboot)
#>
#> Please cite as:
#> Fischer & Roodman. (2021). fwildclusterboot: Fast Wild Cluster.
#> Bootstrap Inference for Linear Regression Models.
#> Available from https://cran.r-project.org/package=fwildclusterboot/.The fwildclusterboot package implements the “fast” wild cluster bootstrap algorithm developed in Roodman et al (2019) for regression objects in R. The “fast” algorithm makes it feasible to calculate test statistics based on a large number of bootstrap draws even for large samples - as long as the number of bootstrapping clusters is not too large.
For linear regression models, fwildclusterboot supports almost all features of Stata’s boottest package. This means that a set of different bootstrap distributions, regression weights, fixed effects, and both restricted (WCR) and unrestricted (WCU) bootstrap inference are supported. The main difference is that it currently only supports univariate hypothesis tests of regression paramters of the form \(H_{0}: R\beta = r\) vs \(H_{1}: R\beta \neq r\), where r is scalar.
Further, fwildclusterboot serves as an R port to WildBootTests.jl, which implements the fast wild cluster bootstrap in Julia at extreme speed. Beyond being significantly faster than fwildclusterboot's native R implementation of the wild cluster bootstrap for OLS (in particular for more demanding problems), WildBootTests.jl offers support for the WRE bootstrap for IV models ((Davidson & MacKinnon, 2010)) and functionality for tests of multiple hypothesis.
A description of the “fast” algorithm is beyond the scope of this vignette. It is very clearly presented in Roodman et al. (2019). For technical details of the implementation in fwildclusterboot, have a look at the technical vignette (tba).
The boottest() function
The fwildclusterboot package consists of one key function, boottest(). It implements the fast wild bootstrap and works with regression objects of type lm, felm and fixest from base R and the lfe and fixest packages.
To start, we create a random data set with two cluster variables (group_id1 & group_id2), two fixed effects and a set of covariates. The icc_ arguments control the cluster variable’s intra-cluster correlation.
# set seed via dqset.seed for engine = "R" & Rademacher, Webb & Normal weights
dqrng::dqset.seed(2352342)
# set 'familiar' seed for all other algorithms and weight types
set.seed(23325)
# load data set voters included in fwildclusterboot
data(voters)
# estimate the regression model via lm
lm_fit <- lm(
proposition_vote ~ treatment + ideology1 + log_income + Q1_immigration ,
data = voters
)
# model with interaction
lm_fit_interact <- lm(
proposition_vote ~ treatment + ideology1 + log_income:Q1_immigration ,
data = voters
)The boottest() function has 4 required and several optional arguments. The required objects are
- object: a regression object of type
lm,fixestorfelm - clustid: a character vector that defines the clustering variables
- param: a character vector of length one - the model parameter to be tested
- B: the number of bootstrap iterations
# boottest on an object of type lm
boot_lm <- boottest(
lm_fit,
clustid = "group_id1",
param = "treatment",
B = 9999
)To tests for an interaction, it is important to use the coefficient names that are internally created by the modeling function.
#names(coef(lm_fit_interact))
boot_lm_interact <- boottest(
lm_fit_interact,
clustid = "group_id1",
param = "log_income:Q1_immigration1",
B = 9999
)boottest() further allows for multivariable tests. Suppose we’re interested in testing the null hypothesis \(0.6*treatment + 0.2*ideology1 = 0.02\). To test such a hypothesis, one would have to specify the hypothesis via the param, R and r arguments:
boot_multi <- boottest(
lm_fit,
clustid = "group_id1",
param = c("treatment", "ideology1"),
R = c(0.6, 0.2),
r = 0.02,
B = 9999
)To access the estimation results, boottest() comes with summary(), tidy() and glance() methods. The tidy() method returns the estimation results in a data.frame. summary() returns additional information on top of the test statistics reported by tidy(). Theglance() method enables the use of output formatting tools from the modelsummary package.
# fwildclusterboot's internal summary() method
summary(boot_lm)
#> boottest.lm(object = lm_fit, param = "treatment", B = 9999, clustid = "group_id1")
#>
#> Hypothesis: 1*treatment = 0
#> Observations: 300
#> Bootstr. Type: rademacher
#> Clustering: 1-way
#> Confidence Sets: 95%
#> Number of Clusters: 40
#>
#> term estimate statistic p.value conf.low conf.high
#> 1 1*treatment = 0 0.073 3.709 0.001 0.033 0.113
summary(boot_multi)
#> boottest.lm(object = lm_fit, param = c("treatment", "ideology1"),
#> B = 9999, clustid = "group_id1", R = c(0.6, 0.2), r = 0.02)
#>
#> Hypothesis: 0.6*treatment+0.2*ideology1 = 0.02
#> Observations: 300
#> Bootstr. Type: rademacher
#> Clustering: 1-way
#> Confidence Sets: 95%
#> Number of Clusters: 40
#>
#> term estimate statistic p.value conf.low
#> 1 0.6*treatment+0.2*ideology1 = 0.02 0.048 2.346 0.025 0.024
#> conf.high
#> 1 0.072
if(requireNamespace("modelsummary")){
# summary via the modelsummary package
library(modelsummary)
msummary(list(boot_lm, boot_lm_interact),
estimate = "{estimate} ({p.value})",
statistic = "[{conf.low}, {conf.high}]")
}
#> Loading required namespace: modelsummary| (1) | (2) | |
|---|---|---|
| 1*treatment = 0 | 0.073 (<0.001) | |
| [0.033, 0.113] | ||
| 1*log_income × Q1_immigration1 = 0 | −0.038 (<0.001) | |
| [−0.057, −0.019] | ||
| Num.Obs. | 300 | 300 |
| R2 | 0.316 | 0.339 |
| R2 Adj. | 0.288 | 0.311 |
| AIC | −82.1 | −92.2 |
| BIC | −30.2 | −40.4 |
| Log.Lik. | 55.025 | 60.102 |
A plot() visualizes bootstrapped p-values as a function of the hypothesized effect size \(r\).
plot(boot_lm)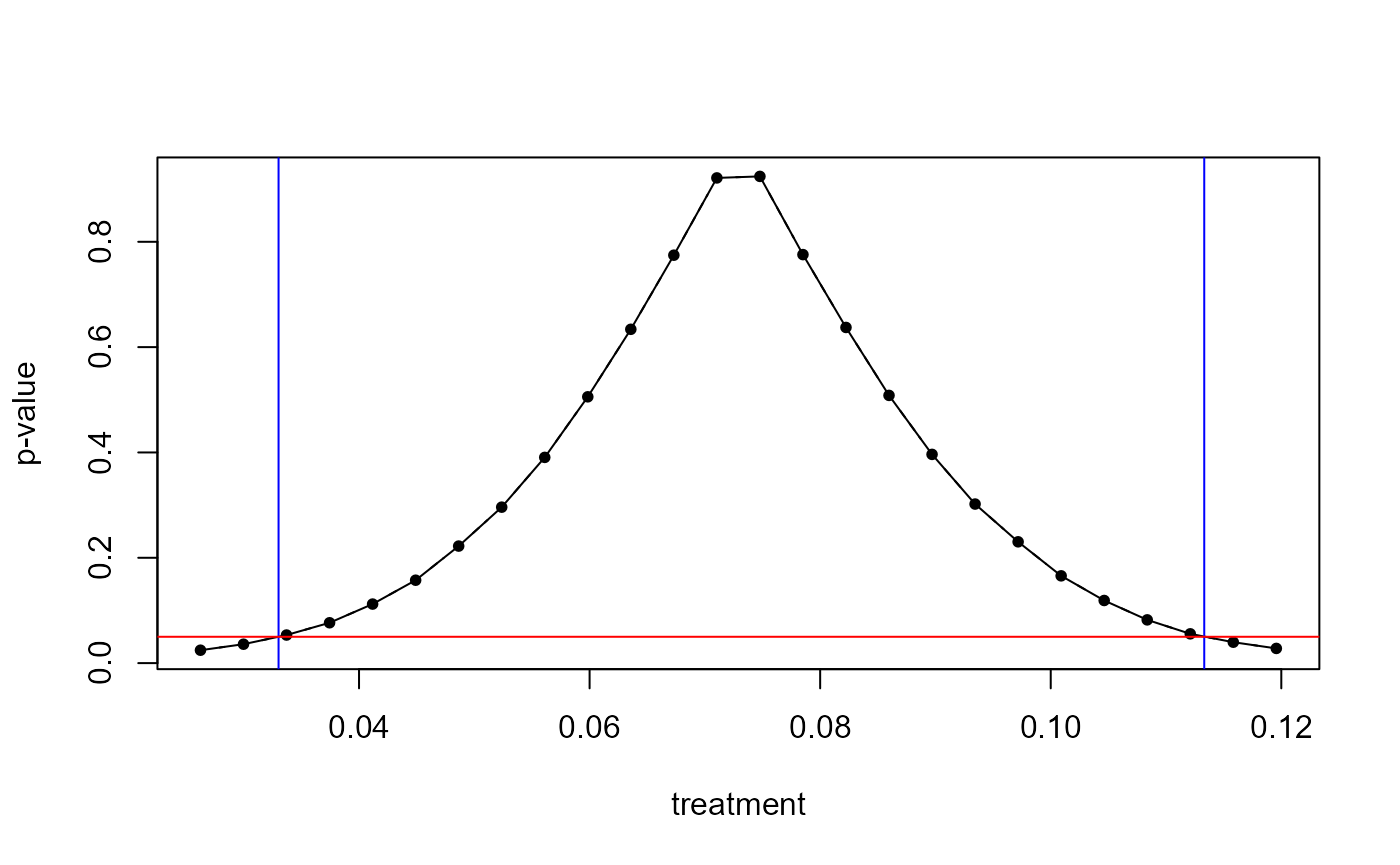
Multiway Clustering
The boottest() function supports clustering of any dimension. E.g. for two-way clustering, one simply needs to specify the names of the cluster variables in a character vector.
boot_lm <- boottest(
lm_fit,
clustid = c("group_id1", "group_id2"),
param = "treatment",
B = 9999
)
summary(boot_lm)
#> boottest.lm(object = lm_fit, param = "treatment", B = 9999, clustid = c("group_id1",
#> "group_id2"))
#>
#> Hypothesis: 1*treatment = 0
#> Observations: 300
#> Bootstr. Type: rademacher
#> Clustering: 2-way
#> Confidence Sets: 95%
#> Number of Clusters: 40 20 253
#>
#> term estimate statistic p.value conf.low conf.high
#> 1 1*treatment = 0 0.073 3.842 0.004 0.031 0.115The Heteroskedastic Bootstrap
If you drop the clustid argument, boottest() will run a heteroskedasticity robust wild bootstrap via the ‘R-lean’ algorithm. At the moment, the null hypothesis is always imposed, only Rademacher and Webb weights are supported, and no confidence intervals are computed. Further, no regression weights are supported. As all algorithms in fwildclusterboot, p-values are calculated based on pivotal t-statistics.
boot_lm <- boottest(
lm_fit,
param = "treatment",
B = 9999
)
summary(boot_lm)
#> boottest.lm(object = lm_fit, param = "treatment", B = 9999)
#>
#> Hypothesis: 1*treatment = 0
#> Observations: 300
#> Bootstr. Type: rademacher
#> Clustering: 0-way
#> Confidence Sets: 95%
#>
#> term estimate statistic p.value conf.low conf.high
#> 1 1*treatment = 0 0.073 3.096 0.002 NA NA
boot_lm$engine
#> [1] "R-lean"Choice of Bootstrap Weights
Furthermore, you can choose among four different weighting distribution via the type argument: Rademacher, Mammen, Normal and Webb. By default, boottest() uses the Rademacher distribution.
boot_lm_rade <- boottest(
lm_fit,
clustid = c("group_id1", "group_id2"),
param = "treatment",
B = 999,
type = "rademacher")
boot_lm_webb <- boottest(
lm_fit,
clustid = c("group_id1", "group_id2"),
param = "treatment",
B = 999,
type = "webb"
)
if(requireNamespace("modelsummary")){
library(modelsummary)
msummary(list(boot_lm_rade, boot_lm_webb),
estimate = "{estimate} ({p.value})",
statistic = "[{conf.low}, {conf.high}]")
}| (1) | (2) | |
|---|---|---|
| 1*treatment = 0 | 0.073 (0.006) | 0.073 (0.006) |
| [0.031, 0.115] | [0.029, 0.117] | |
| Num.Obs. | 300 | 300 |
| R2 | 0.316 | 0.316 |
| R2 Adj. | 0.288 | 0.288 |
| AIC | −82.1 | −82.1 |
| BIC | −30.2 | −30.2 |
| Log.Lik. | 55.025 | 55.025 |
Other function arguments
Via the function argument sign_level, you can control the significance level of the test. The default value is sign_level = 0.05, which corresponds to a 95% confindence interval.
boot_lm_5 <- boottest(
lm_fit,
clustid = c("group_id1"),
param = "treatment", B = 9999,
sign_level = 0.05
)
boot_lm_10 <- boottest(
lm_fit,
clustid = c("group_id1"),
param = "treatment", B = 9999,
sign_level = 0.10
)
if(requireNamespace("modelsummary")){
library(modelsummary)
msummary(list(boot_lm_5, boot_lm_10),
estimate = "{estimate} ({p.value})",
statistic = "[{conf.low}, {conf.high}]")
}| (1) | (2) | |
|---|---|---|
| 1*treatment = 0 | 0.073 (<0.001) | 0.073 (<0.001) |
| [0.033, 0.113] | [0.040, 0.106] | |
| Num.Obs. | 300 | 300 |
| R2 | 0.316 | 0.316 |
| R2 Adj. | 0.288 | 0.288 |
| AIC | −82.1 | −82.1 |
| BIC | −30.2 | −30.2 |
| Log.Lik. | 55.025 | 55.025 |
In the case of multiway clustering, one might want to specify the bootstrap clustering level. By default, boottest chooses the clustering level with the highest number of clusters as bootcluster = "max". Other choices are the minimum cluster, or independent clustering variables.
boot_lm1 <- boottest(
lm_fit,
clustid = c("group_id1", "group_id2"),
param = "treatment",
B = 9999,
bootcluster = "min"
)
boot_lm2 <- boottest(
lm_fit,
clustid = c("group_id1", "group_id2"),
param = "treatment",
B = 9999,
bootcluster = "group_id1"
)
if(requireNamespace("modelsummary")){
library(modelsummary)
msummary(list(boot_lm1, boot_lm2),
estimate = "{estimate} ({p.value})",
statistic = "[{conf.low}, {conf.high}]")
}| (1) | (2) | |
|---|---|---|
| 1*treatment = 0 | 0.073 (0.005) | 0.073 (0.008) |
| [0.031, 0.111] | [0.027, 0.115] | |
| Num.Obs. | 300 | 300 |
| R2 | 0.316 | 0.316 |
| R2 Adj. | 0.288 | 0.288 |
| AIC | −82.1 | −82.1 |
| BIC | −30.2 | −30.2 |
| Log.Lik. | 55.025 | 55.025 |
Fixed Effects
Last, boottest() supports out-projection of fixed effects in the estimation stage via lfe::felm() and fixest::feols(). Within the bootstrap, it is possible to project out only one fixed effect, which can be set via the fe function argument. All other fixed effects specified in either felm() or feols() are treated as sets of binary regressors. Note that boottest.fixest() currently does not know how to properly handle advanced fixest syntax for fixed effects, as e.g. varying slopes.
if(requireNamespace("fixest")){
# estimate the regression model via feols
library(fixest)
feols_fit <- feols(
proposition_vote ~ treatment + ideology1 + log_income |
Q1_immigration ,
data = voters
)
boot_feols <- boottest(
feols_fit,
clustid = "group_id1",
param = "treatment",
B = 9999,
fe = "Q1_immigration"
)
}
#> Loading required namespace: fixest
#> Too guarantee reproducibility, don't forget to set a global random seed
#> **both** via `set.seed()` and `dqrng::dqset.seed()`.
#> This message is displayed once every 8 hours.The Subcluster Bootstrap
In the case of few treated clusters, MacKinnon and Webb (2018) suggest to use subclusters to form the bootstrap distribution. boottest() allows to specify subclusters via the bootcluster argument.
boot_min <- boottest(
lm_fit,
clustid = c("group_id1", "group_id2"),
param = "treatment",
B = 9999,
bootcluster = "min"
)
boot_var <- boottest(
lm_fit,
clustid = c("group_id1", "group_id2"),
param = "treatment",
B = 9999,
bootcluster = "group_id1"
)
boot_2var <- boottest(
lm_fit,
clustid = c("group_id1", "group_id2"),
param = "treatment",
B = 9999,
bootcluster = c("group_id1", "Q1_immigration")
)
if(requireNamespace("modelsummary")){
library(modelsummary)
msummary(model = list(boot_min, boot_2var),
estimate = "{estimate} ({p.value})",
statistic = "[{conf.low}, {conf.high}]")
}| (1) | (2) | |
|---|---|---|
| 1*treatment = 0 | 0.073 (0.003) | 0.073 (0.010) |
| [0.032, 0.110] | [0.027, 0.119] | |
| Num.Obs. | 300 | 300 |
| R2 | 0.316 | 0.316 |
| R2 Adj. | 0.288 | 0.288 |
| AIC | −82.1 | −82.1 |
| BIC | −30.2 | −30.2 |
| Log.Lik. | 55.025 | 55.025 |
Regression Weights / Weighted Least Squares (WLS)
If regression weights are specified in the estimation stage via lm(), feols() or felm(), boottest() incorporates the weights into the bootstrap inference:
Parallel Execution
A major bottleneck for the performance of boottest() is a large matrix multiplication, which includes the bootstrap weights matrix on the right. In order to speed up the computation, this multiplication calls the c++ Eigen library, which allows for parallelization of dense matrix products. By default, boottest() uses one thread. Note that there is a cost of parallelization due to communication overhead. As a rule of thumb, if boottest() takes more than 10 seconds per execution, using a second thread might speed up the bootstrap.
The number of threads can be specified via the nthreads argument of boottest():
boot_lm <- boottest(
lm_fit,
clustid = "group_id1",
param = "treatment",
B = 9999,
nthreads = 2
)Running the wild cluster bootstrap via WildBootTests.jl
fwildclusterboot serves as an R port to the WildBootTests.jl package.
For guidance on how to install Julia and WildBootTests.jl and how to connect R and Julia, please take a look at the running WildBootTests.jl through fwildclusterboot vignette.
You can tell boottest() to run WildBootTests.jl by using the engine function argument:
boot_lm <- boottest(
lm_fit,
clustid = "group_id1",
param = "treatment",
B = 9999,
engine = "WildBootTests.jl"
)
tidy(boot_lm)
# term estimate statistic p.value conf.low conf.high
#1 1*treatment = 0 0.07290769 3.709435 0.00060006 0.03326969 0.1134117The seed used within Julia is linked to R’s global seed, which you can set through the familiar set.seed() function.
If you decide to run all your bootstraps through WildBootTests.jl, you can set a global variable via
setBoottest_engine("WildBootTests.jl")Calling boottest() without specifying engine = "WildBootTests.jl" will now automatically run the bootstrap through WildBootTests.jl.
The WRE bootstrap for IV models
Through WildBootTests.jl, fwildclusterboot supports the WRE bootstrap by Davidson & MacKinnon, 2010 for IV (instrumental variables) models for objects of type ivreg via the boottest() function:
library(ivreg)
data("SchoolingReturns", package = "ivreg")
# drop all NA values from SchoolingReturns
SchoolingReturns <- na.omit(SchoolingReturns)
ivreg_fit <- ivreg(
log(wage) ~ education + age + ethnicity + smsa + south + parents14 |
nearcollege + age + ethnicity + smsa + south + parents14,
data = SchoolingReturns)
boot_ivreg <- boottest(
object = ivreg_fit,
B = 999,
param = "education",
clustid = "kww",
type = "mammen",
impose_null = TRUE
)
tidy(boot_ivreg)
# term estimate statistic p.value conf.low conf.high
# 1 1*education = 0 0.0638822 1.043969 0.2482482 -0.03152655 0.2128746Tests of multiple joint hypotheses (q > 1)
Through WildBootTests.jl, you can also test multiple joint hypotheses via the mboottest() function. With minor differences, mboottest()'s syntax largely mirrors boottest().
To jointly test the null hypothesis \(H_0: treatment = 0 \text{ and } ideology1 = 0\) vs \(H_0: treatment \neq 0 \text{ and } ideology1 \neq 0\) via a wild cluster bootstrap, you can run
library(clubSandwich)
R <- clubSandwich::constrain_zero(2:3, coef(lm_fit))
wboottest <-
mboottest(
object = lm_fit,
clustid = "group_id1",
B = 999,
R = R
)
tidy(wboottest)
# teststat p_val
# 1 8.469086 0Miscellanea
A sanity check if fwildclusterboot::boottest() works as intended
A sanity check to see if fwildclusterboot::boottest() works as intended is to look at its t_stat return value. For both the WCR and WCU bootstrap, boottest() re-calculates the “original” - hence non-bootstrapped - t-statistic from its input regression model. The t-stat computed in boottest() and the t-stats reported by either lm(), feols() and lfe() under the same error clustering structure and small-sample adjustments should be identical. If you find that they differ, please report a bug on github. Note that fwildclusterboot explicitly tests for t-stat equality against fixest::feols() here.
data <-
fwildclusterboot:::create_data(
N = 1000,
N_G1 = 20,
icc1 = 0.81,
N_G2 = 10,
icc2 = 0.01,
numb_fe1 = 10,
numb_fe2 = 10,
seed = 8769
)
# oneway clustering
feols_fit <- fixest::feols(
proposition_vote ~ treatment + ideology1 + log_income,
data = data,
cluster = ~group_id1,
ssc = fixest::ssc(adj = TRUE,
cluster.adj = TRUE,
cluster.df = 'conventional')
)
feols_tstats <- fixest::coeftable(
feols_fit
)[c("treatment", "log_income", "ideology1"), 3]
boot_tstats <-
lapply(c("treatment", "log_income", "ideology1"), function(x){
boot1 <- fwildclusterboot::boottest(
feols_fit,
clustid = c("group_id1"),
B = 999,
param = x,
ssc = fwildclusterboot::boot_ssc(
adj = TRUE,
cluster.adj = TRUE,
cluster.df = 'conventional'),
impose_null = TRUE)$t_stat
})
df <- cbind(feols_tstats, unlist(boot_tstats))
colnames(df) <- c("feols tstat", "boottest tstat")
df
#> feols tstat boottest tstat
#> treatment 0.9958071 0.9958071
#> log_income -2.9129869 -2.9129869
#> ideology1 1.4169933 1.4169933Small Sample Corrections
boottest() offers several options for small sample adjustments via the ssc function argument which need to be specified via the boot_ssc() function. boot_ssc() has 4 arguments and is intentionally designed to mimic fixest's ssc() function. For more information on the default choices and alternative options, see ?fwildclusterboot::boot_ssc().
Memory & the ‘lean’ implementation of the wild cluster bootstrap
Because the R-implementation of the fast algorithm is memory-intensive, fwildclusterboot further supports a Rcpp-based ‘lean’ implementation of the wild cluster bootstrap in case that memory demands get prohibitively large. In general, the ‘lean’ algorithm is much slower: its main feature is that it requires much less memory. The algorithm is equivalent to the ‘wild2’ algorithm in the “Fast & Wild” paper by Roodman et al. Note that the implementation in WildBootTests.jl is, in general, very memory-efficient.
library(bench)
dt <- fwildclusterboot:::create_data(
N = 10000,
N_G1 = 250,
icc1 = 0.01,
N_G2 = 10,
icc2 = 0.01,
numb_fe1 = 10,
numb_fe2 = 10,
seed = 7645
)
lm_fit <- lm(
proposition_vote ~ treatment + ideology1 + log_income + Q1_immigration ,
data = dt
)
res <-
bench::mark(
"R" = boottest(lm_fit,
clustid = "group_id1",
param = "treatment",
B = 9999,
engine = "R",
nthreads = 4),
"R-lean" = boottest(lm_fit,
clustid = "group_id1",
param = "treatment",
B = 9999,
engine = "R-lean",
nthreads = 4),
"WildBootTests.jl" =
boottest(lm_fit,
clustid = "group_id1",
param = "treatment",
B = 9999,
engine = "WildBootTests.jl"),
iterations = 1,
check = FALSE
)
resSeeds
To guarantee reproducibility, you need to set a global random seed via
-
set.seed()when using- the lean algorithm (via
engine = "R-lean"), - the heteroskedastic wild bootstrap
- the wild cluster bootstrap via
engine = "R"with Mammen weights or engine = "WildBootTests.jl"
- the lean algorithm (via
-
dqrng::dqset.seed()when usingengine = "R"for Rademacher, Webb or Normal weights
Treatment of Invalid Test Statistics for multiway clustering
In case of multi-way clustering, it is not guaranteed that the covariance matrix is positive definite, in which case the resulting bootstrap test statistics are invalid. boottest() follows the implementation in STATA and deletes invalid tests statistics, and informs the user with a note.
